TRANSPOSONS, PLASMIDS AND BACTERIOPHAGES:
Transposons
Transposons are sequences of DNA that can move or transpose themselves to new positions within the genome of a single cell.
They are also known as 'jumping genes'.
Transposition can create phenotypically significant mutations and alter the cell's genome size.
- The first transposons were discovered in maize {Zea mays), by Barbara McClintock in 1948, for which she was awarded a Nobel Prize in 1983.
- About 50% of the total genome of maize consists of transposons.
- In bacteria, transposons can jump from chromosomal DNA to plasmid DNA and back.
- It leads to the transfer and permanent addition of genes such as those encoding for antibiotic resistance.
- Multi-antibiotic resistant bacterial strains can be generated in this way.
- In primates, including humans, some sequences of interspersed repetitive DNA are present.
- This common form of transposon in humans is called 'Alu' sequence. It is approximately 300 bases long.
On the basis of their mechanism they are classified into two types;
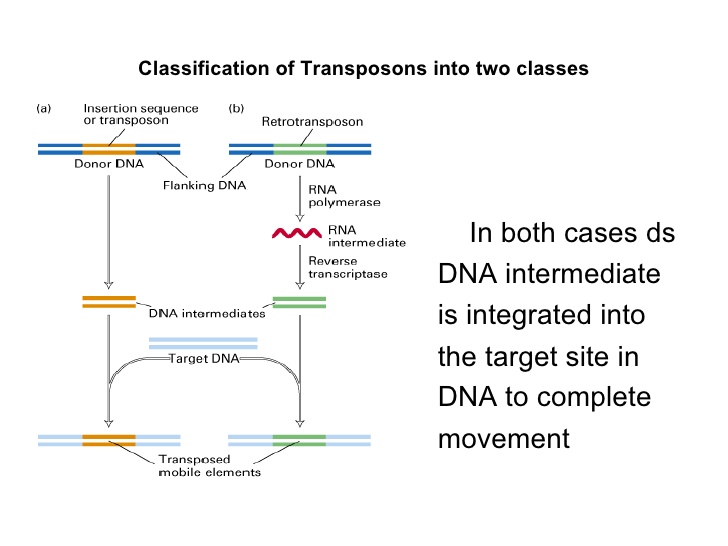
Retrotransposons:
They copy in two stages, first from DNA to RNA by transcription and then from RNA back to DNA by reverse transcription.
The DNA copy is then inserted in the new posiion. ("Copy and paste")
DNA transposons:
They do not involve RNA intermediate.
The enzyme transposase makes a staggered cut at the target site producing sticky ends, cuts out the transposon and ligates in new position.
("Cut and paste")
Plasmids
Plasmids are the small, extra-chromosomal,'double stranded, circular forms of DNA that replicate autonomously.
Most of the plasmids are not required for the survival of the organism in which they reside.
They are naturally found in bacteria, yeast and occasionally in plant and animal cells.
The term plasmid was first introduced by the American molecular biologist Joshua Lederberg in 1952.
Plasmid size varies from 1 to over as 1,000 kilo base pairs (kbp).
Plasmids are considered as transferable genetic elements, or 'replicons', capable of autonomous replication within a suitable host.
Plasmids in bacteria carry genes related to metabolic activity and allowing the carrier bacteium to survive and reproduce under unfavourable conditions.
Plasmids cannot accomodate long DNA fragments therefore cosmids are constructed.
Cosmids are plasmids having fragments of lambda phage vectors.
Nomenclature of Plasmids
It is a common practice to designate plasmid by
1. a lower case P (p)
2. the first letter/letters of researcher(s) names
3. the numerical number given by the workers.
Thus, pBR 322 is as
1. a lower case P (p)
2. plasmid discovered by Bolivar and Rodriguez,
3. designated it as 322.
Some plasmids are given names of the places where they are discovered
e.g. pUC is plasmid from University of California.
Bacteriophage
A bacteriophage (Greek 'phagein': to eat) is a virus that infects bacteria.
Commonly its short form, 'phage' is used.
Typically, bacteriophages consist of an outer protein capsid enclosing the genetic material.
The 'genetic material can be ssRNA, dsRNA, ssDNA, or dsDNA ('ss-' or 'ds-' prefix denotes single-strand or double-strand) in either circular or linear arrangement.
Bacteriophages are much smaller than the bacteria they destroy.
The cloning of single genes is usually best .carried out using plasmid.
However, for cloning of larger pieces of DNA (i e. during gene library construction) these plasmids are not suitable as larger inserts increase the plasmid size making the transformation inefficient.
.Larger DNA molecules can be injected in host bacterial cell by bacteriophages.
Commonly used bacteriophages as cloning vectors are M13 and lambda-phage which infect E.coli.
Phage λ or Lambda phage
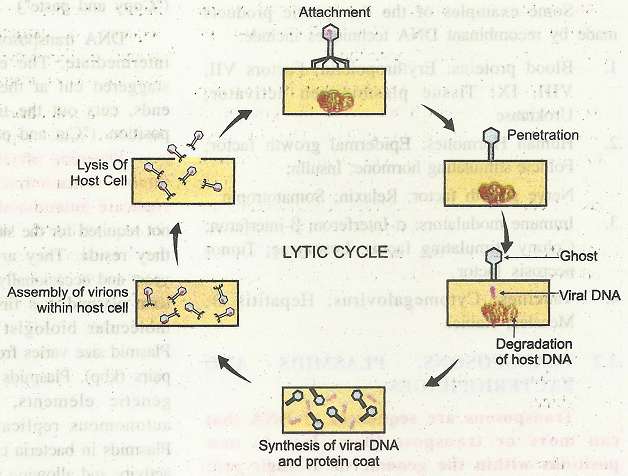
The DNA size of phage is 48.5 kb in length. It is with cos sites of 12 bp at the ends. The cohesive cos ends allow the DNA to be circularized in the host cell. For the cloning of large DNA fragments up to about 20 kbp DNA of phage is removed. It is replaced by the DNA with desired gene. The recombinant DNA is then packaged within viral particles in vitro. These are allowed to infect bacterial cells which have been cultured on agar. 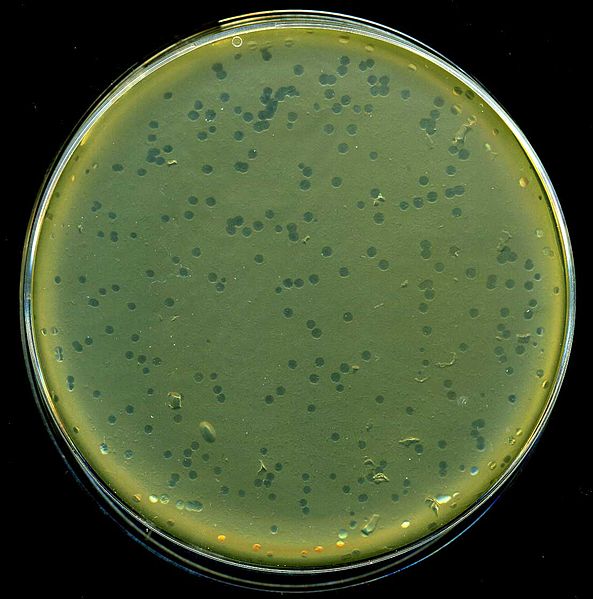 Once inside the bacterial cells, the recombinant viral DNA starts replicating. All the genes needed for multiplication of the virus and normal lytic cycle are still present in the DNA. Growth takes place by cycles of cell lysis and the released viruses infect the surrounding bacterial cells. It gives rise to plaques of lysed bacterial cells on a background or, lawn of bacterial cells. Cloned DNA can be recovered from the viruses in these plaques. Replication of bacteriophages, (lytic cycle) inside the specific host/ bacterial cell takes place in following steps:
Once inside the bacterial cells, the recombinant viral DNA starts replicating. All the genes needed for multiplication of the virus and normal lytic cycle are still present in the DNA. Growth takes place by cycles of cell lysis and the released viruses infect the surrounding bacterial cells. It gives rise to plaques of lysed bacterial cells on a background or, lawn of bacterial cells. Cloned DNA can be recovered from the viruses in these plaques. Replication of bacteriophages, (lytic cycle) inside the specific host/ bacterial cell takes place in following steps:
1. Attachment: Bacteriophages attach to specific receptors on the surface of bacteria. As phages do not move independently, they rely on random encounters with the right receptors.
2. Penetration: After the contact, the tail fibres bring the base plate closer to the surface of the cell. Once attached completely, the tail contracts, injecting genetic material (DNA) through-'bacterial membrane. (Capsid -protein coat remains outside and is called 'ghost')
3. Synthesis of proteins and nucleic acids: The host's normal synthesis of proteins and nucleic acids is disrupted, and it is forced to manufacture viral DNA, and proteins instead. These products are the parts of new virions within the ceil or proteins involved in cell lysis.
4. Virion assembly: The base plates are assembled with the tails first. The heads-capsids are constructed separately and then are joined with the tails. The DNA is packed efficiently within the heads. The whole process takes about 15 minutes.
5. Release of virions: Phages are released via lysis of cell. It is achieved by an enzyme called endolysin, which breaks down the cell wall. Released virions are capable of infecting a new bacterium.