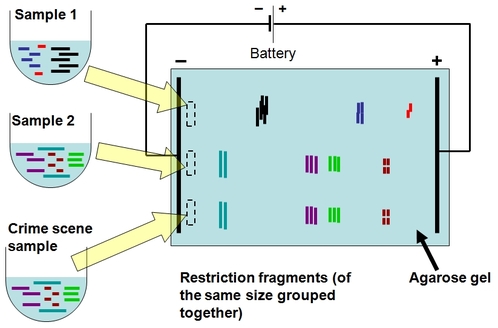
RESTRICTION FRAGMENTS:
A restriction fragment is a DNA fragment resulting from the cutting of a DNA strand by the reslriction enzyme (restriction endonucleases orRENs), a process called restiction.
Steward Linn and Werner Arber isolated two enzymes in 1963 which restricted the growth of bacteiophage in bacteium E. coli.
One of these enzymes cut DNA and was named restriction endonuclease.
Restriction enzymes belong to a larger class of enzymes called nucleases.
The nucleases are of two types;
1 .Exonucleases- They remove nucleotides from the ends of the DNA.
2. Endonucleases- They make cuts at specific positionswithin the DNA.
Restriction endonucleases serve as tools for cutting DNA molecules at predetermined sites which is the basic requirement for gene cloning or recombinant DNA technology.
Types of REN:
- Three main types- type I, type II, and type III are known.
- They have slightly different modes of action.
- Type II restriction enzymes are used in rDNA technology because they can be used in vitro to identify and cleave within specific DNA sequences usually having 4-8 nucleotides.
- More than 350 different type II endonucleases with 100 different recognition sequences, are. known.
Nomenclature:
Restriction endonucleases are named by a standard procedure, with particular reference io the bacteria from which they are isolated.
1. The first letter (in italics) of the enzymes indicates the genus name
2. The first two letters (also in italics) of the species,
3. the strain of the organism
4. a Roman numeral indicating the order of discovery.
A couple of examples are given below:
EcoRI is from Escherichia (E) coli (co) strain Ry 13 (R) and first endonuclease (I) to be discovered.
Hind III is from Haemophilus (H) influenzae (in) strain Rd (d) and the third endoculease (III) to be discovered.
Recognition Sequences:
Recognition sequence or restiction site is the site where the DNA is cut by a restriction endonuclease.
Restriction endonucleases speciically recognize DNA with a particular sequence of 4-8 nucleotides and cleave.
Each restriction enzyme is highly specific, recognizing a particular short DNA sequence, or restriction site, and cutting both DNA strands at the same time at specific points within that site.
Most restriction sites are palindromes.
In a palindrome, base sequence of second half in DNA strand represents the mirror image of the base sequence of first half.
Palindromes are groups of letters that form the same word when read both forward and backward,
MALAYALAM word is reads same backward and forward in a palindrome word.
Palindrome in DNA is a sequence of base pairs that reads the same on the two strands when orientation of reading is kept the same.

Cleavage Patterns:
In recombinant DNA technology specific restriction endonucleases (particularly type II).
It recognize a specific base pair in DNA called a restriction site.
It cleave the DNA within the sequence by hydrolyzing the phosphodiester bonds(retaining symmetry).
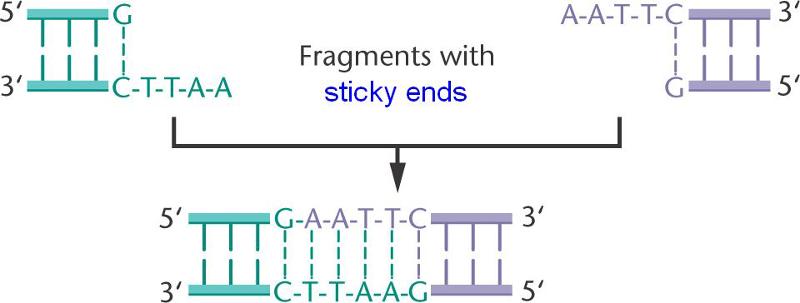
Hence the double stranded restriction fragments have single stranded ends.
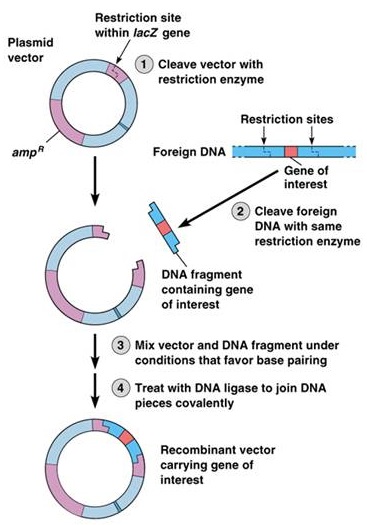
These short extensions, called 'sticky ends' can form hydrogen bonded base pairs with complementary sticky ends on any other DNA cut with the same enzyme (such as a bacterial plasmid).
They require Mg++ ions for cleavage.
Many cuts are made by one restriction enzyme because of the chances of repetition of these sequences in a long DNA molecule, yielding a set of restriction fragments.
A particular DNA molecule will always yield the same set of restriction fragments when exposed to the same restriction enzyme.
Restriction fragments can be analyzed using techniques such as gel electrophoresis or used in recombinant DNA technology.
In agarose gel electrophoresis, the restriction fragments yield a band pattern characteristic of the original DNA molecule and restriction enzyme used,
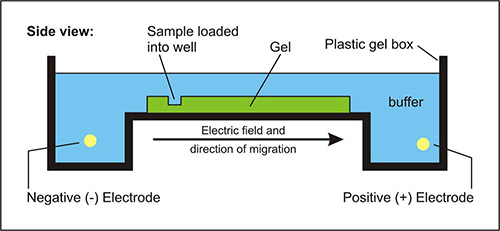
for example, the relatively small DNA molecules of viruses and plasmids can be identified simply by their restriction fragment patterns.